Welcome
Welcome to the Joseph Group at Princeton University. We are part of the Department of Chemical and Biological Engineering and the Omenn-Darling Bioengineering Institute.

Biomolecular condensates
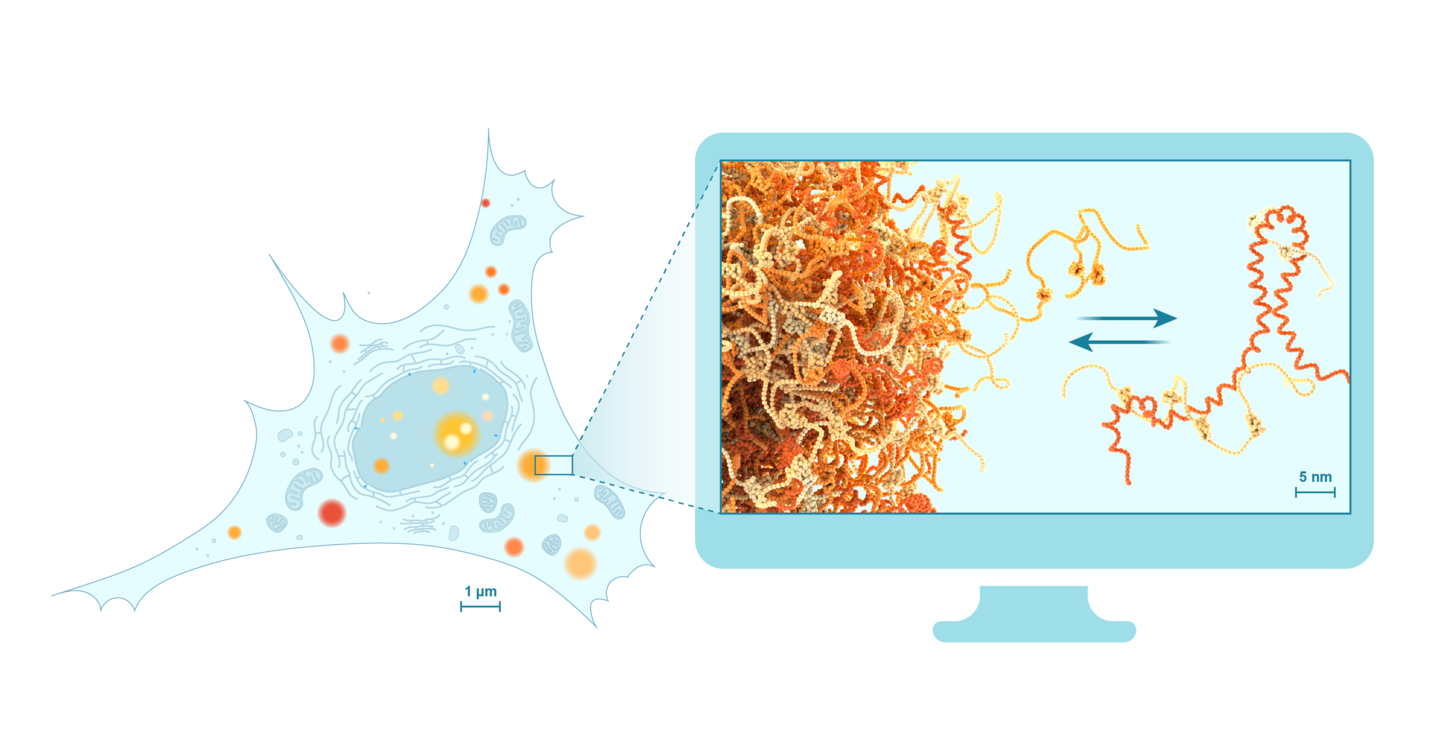
We focus on the characterization of biomolecular condensates—liquid-like organelles inside cells that lack physical membranes. These structures are largely composed of proteins, RNA, and other biomolecules and are thought to assemble via phase separation.
Importantly, biomolecular condensates are linked to both normal cellular functions (e.g., RNA processing, stress adaptation, signaling) and cellular dysfunction (e.g., neurodegenerative disorders, cancers, infectious diseases). Thus, there is an urgent need to understand how the material properties of condensates are encoded and how these give rise to complex cellular functions and disease.
Decoding the design rules of condensates will allow us to engineer therapeutics for targeting condensate-linked diseases. Additionally, due to their dynamic liquid-like nature and lack of membranes, biomolecular condensates present exciting opportunities for engineering new cellular functions (e.g., designing condensates that act as microreactors, biosensors, etc.).
Our research focuses on designing chemically-specific models of condensates to zoom into these structures and understand how interactions of proteins and RNA give rise to emergent condensate behaviors. Additionally, we leverage our models to accelerate condensate engineering.
Key techniques and approaches
We specialize in developing chemically-specific coarse-grained models of proteins and RNA, enabling the chemically specific recapitulation of condensates and other biomolecular structures.
Our work integrates diverse approaches and techniques, including atomistic simulations, coarse-grained modeling, molecular dynamics, Monte Carlo sampling, machine learning, and statistical mechanics.
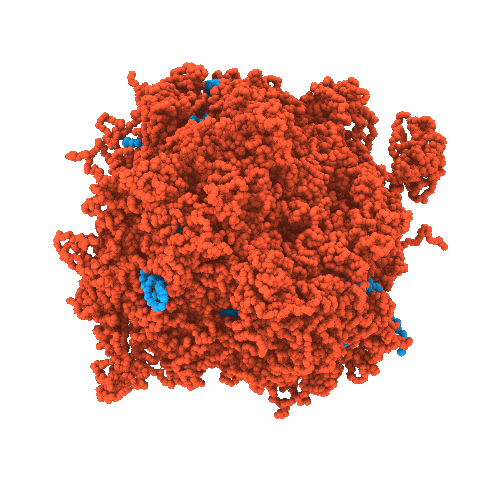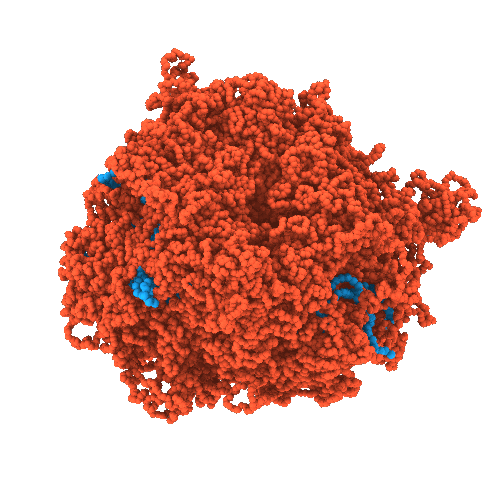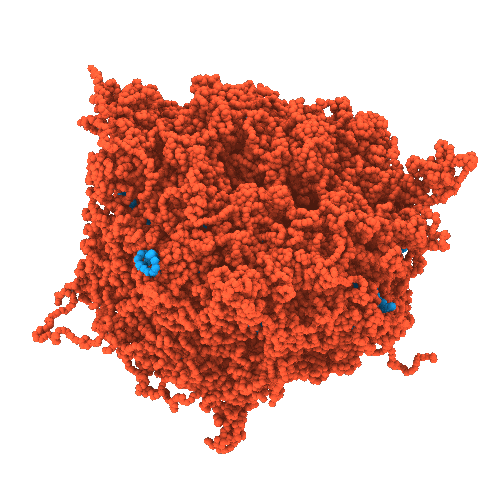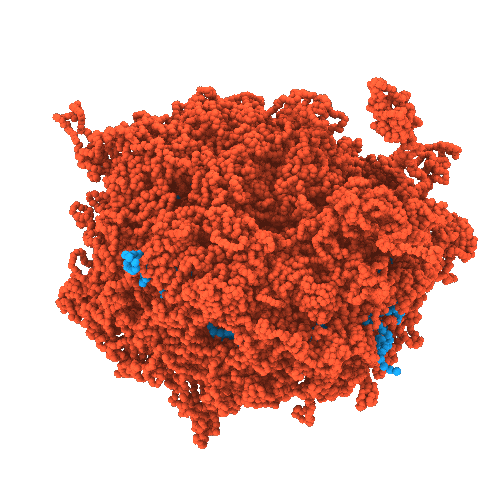
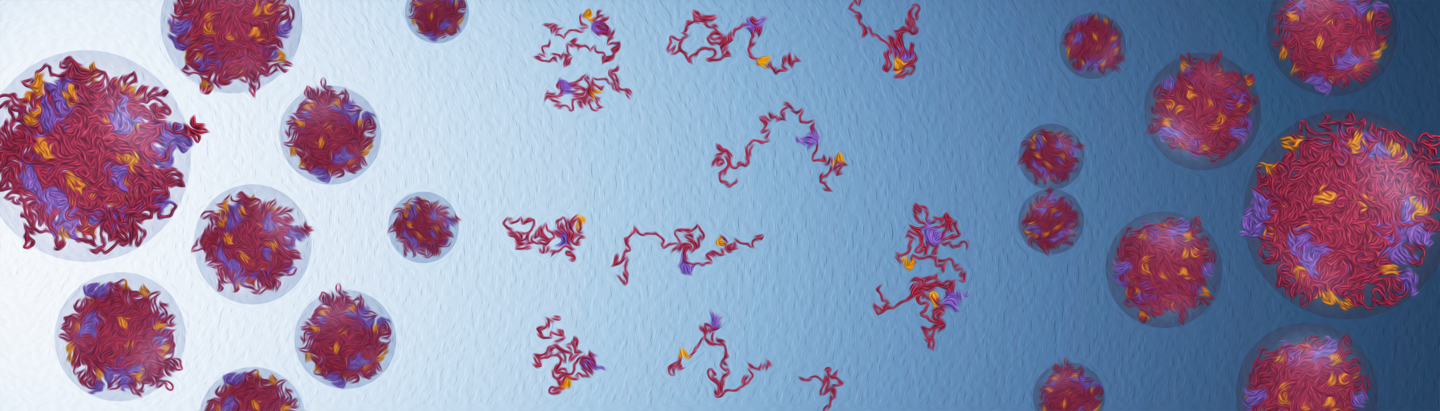
Developing chemically-specific models of condensates

- Parameterization of protein and RNA models
- High-resolution structural characterization of condensates
- Prediction of thermodynamic and transport properties
Characterizing condensates for biomedical applications

- Physical determinants of condensate maturation and aging
- Interactions between small molecules (e.g., drugs) and condensates
- Designing condensate modulators (e.g., molecules that disrupt condensates)
Engineering condensates for sustainability applications

- Designing condensates from protein and RNA sequence
- Engineering enzymatic condensates for natural product synthesis (metabolic engineering)